Challenges in implementing genomic medicine: the 100,000 Genomes Project
Abstract
Many important medical conditions may be the result of an inherited mutation in one of a number of different genes. Technical advances have reduced the cost of whole genome sequencing and whole exome sequencing to a level where it is now feasible to analyse multiple genes in one test. Every human carries several hundred potentially pathogenic coding variants, so a major challenge is to understand which of these is relevant to the patient’s disease. This requires considerable computing power, the use of international unaffected “normal” population and disease cohort databases, clinical scientist input as well as a medical context provided by accurate phenotyping and understanding of Bayesian probability. The guidelines of the American College of Medical Genetics provide a useful framework for the evaluation and reporting of sequence variants. In England, the 100,000 Genomes Project was established within the National Health Service to introduce these technologies into mainstream healthcare. From January 2019, genomic laboratory hubs will deliver a genomics service. In this paper, we review what has been learnt from the project to date and consider how other health care systems could use similar approaches.
Keywords
Introduction
Genomic medicine
Given the brief of this review to assess the impact of the 100,000 Genomes Project on clinical care, we define genomic medicine as the diagnosis, prediction, prognosis, prevention and/or treatment of disease and disorders of the mind and body, using approaches informed or enabled by knowledge of the genome and the molecules it encodes. In this paper, we focus on the use of DNA sequence data to aid in the diagnosis and planning the initial management of cancer and rare inherited disorders, which often have a mendelian inheritance pattern. There is a wider role of genomic medicine in population risk assessment, prognostic assessment, companion tests [e.g., Estrogen receptor (ER) status in breast cancer] or comprehensive treatment decision aids (e.g., oncotype diagnosis for breast cancer) and therapy monitoring (e.g., in acute leukaemias) which are not discussed in detail.
The UK Clinical Genetics services
The following case history illustrates the sequence of events for a typical referral in the “pre-genomics” era. We will go on to describe how the 100,000 Genomes Project is being used as a catalyst within the NHS to introduce genomic medicine with linked diagnostics and therapeutics.
A 34-year-old lady presents to her primary care physician (general practitioner) in the UK NHS with a breast lump. She is referred to her district general hospital breast (secondary) care team for an examination, scan and a biopsy. A cancer diagnosis is made, and a date for surgery is set. A family history is taken, when it is established that her paternal aunt was diagnosed with breast cancer under the age of forty. The patient is referred to a specialist clinical genetics (tertiary care) service. She sees a counsellor to discuss the pros and cons of having a BRCA1 and BRCA2 gene test. Germline genetic testing of a blood sample is undertaken in the regional molecular laboratory. This demonstrates a pathogenic alteration in BRCA1 which is reported a month post initial surgery. The clinical genetics department informs the patient, surgeon and the oncologist to help plan if further preventative surgery is recommended. A cascade letter is written for relatives to access predictive testing, screening and, when appropriate, reproductive counselling and/or risk-reducing surgery. The patient is informed of relevant patient support groups and offered additional psychological support if required.
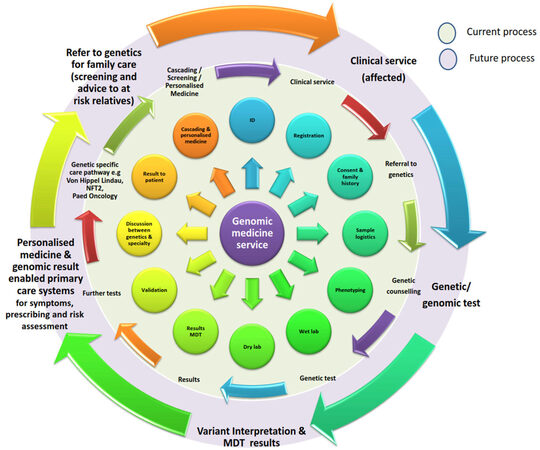
In the near future, UK pathways for cancer diagnosis, or the investigation of rare Mendelian disorders will change radically [Figure 1]. Much diagnostic genetic testing will move from clinical genetics departments. Instead, it will be initiated by clinicians in the relevant specialty; this is called “mainstreaming”. Whole genome sequencing (WGS) and whole exome sequencing will be done in consolidated laboratories. In an attempt to improve our understanding and interpretation of genomic variation, phenotypic data will be collected about patients to assist in the assessment of their genomic results. These approaches are a direct legacy of the 100,000 Genomes Project.
The 100,000 Genomes Project
In 2012, the then Prime Minister David Cameron announced funding for the 100,000 Genomes Project, to be organised by Genomics England (GE), a private company formed by the Department of Health in 2013. Through this project, GE works in partnership with NHS England (itself a non-departmental public body of the Department of Health) to integrate WGS into the NHS. The 100,000 Genomes Project aims to sequence 100,000 genomes from NHS patients with cancer and rare diseases. Data collected from the 100,000 Genomes Project can inform research on rare diseases, or benefit patient care potentially by streamlining the diagnostic process and tailoring care to the individual.
The project has strict inclusion criteria, to ensure data of clinical and research benefit is gathered. For over 300 rare diseases, specific criteria[1] are applied to maximise chance of recruiting individuals whose disease may have a Mendelian basis. The project also requires submission of phenotypic data using SNOMED[2] (standardised healthcare terminology used for electronic health records and coding in over fifty countries https://www.snomed.org/snomed-ct/what-is-snomed-ct) terms, and evidence of previous genetic testing (to screen out previously known mutations). A patient’s whole blood samples must pass quality assurance and quality control tests. When relevant, close relatives (usually the parents) of the patient also undergo WGS. In the case of autosomal dominant conditions all affected members may be sequenced.
Referrals of qualifying patients are sent by clinicians to one of thirteen genomic medicine centres across the country where samples are collected. For all patients this involves a whole blood sample, but for cancer patients, an additional requirement is a fresh frozen sample from the tumour, to provide a contrasting genome to the germline. Tissue samples must yield high quality DNA, as formalin-fixed paraffin-embedded samples rarely provide high quality DNA for sequencing. These must be processed within 24 h to avoid denaturation of the DNA.
After storage and checks at the UK Biocentre, samples are sent to a NHS genomic sequencing centre for sequencing and tier analysis using a crowd-sourced database called PanelApp[3]. This database lists genes that are potentially involved in each rare disease studied by the project. Experts worldwide can add to this list of genes and review the degree of evidence supporting their involvement in these conditions. The evidence for each gene is ranked with a traffic light system from green (meaning high diagnostic-grade level of evidence that the gene is involved) to red (low evidence). This provides a global consensus, which helps to standardise the genes tested for each disease. After an expert panel from GE has evaluated the gene panel, it can be used to interpret genomes from the project. The number of genes added to these panels is expected to increase as researchers learn from the data collected by the 100,000 Genomes Project. The analysis sorts results into five categories: nonsense or frameshift in known genes, missense in known genes, frameshift in suspected genes, carrier findings and additional incidental findings. Results are then confirmed and collated at the data centre.
At the data centre, the resulting data are made available both as anonymised research data, as well as combined with identifiable data to provide clinicians with practical conclusions from the sequencing analysis. This clinical data is also accessible for research purposes through a coalition of NHS and academic researchers that form the GE Clinical Interpretation Partnership (GeCIP), which itself is split into 42 domains based on speciality.
Patient engagement
The 100,000 Genomes Project has sought to catalyse the development of a full genomic medicine service through variant interpretation, gene mapping and the provision of information about the effectiveness of treatment based on inherited and somatic variants. In addition the project has included initiatives to engage patients in the development of a genomic medicine service[4]. This is to reassure patients about the data sharing and confidentiality agreements. This is important given the aim to generate a clinical, research and pharmaceutical industry facing matched and anonymised clinical and genomic record database that can be linked to other international cohorts.
We have developed a stakeholder group with both individual patient and established support group representation for outreach, oversight, insight, lobbying and individual patient support. This has resulted in a number of projects to engage seldom heard from communities, a prominent national media campaign, and leading a national helpline. In addition we have contributed evidence to two parliamentary select committee[5,6] reports that have changed national policy on the role of the National Health Service (NHS) UK-wide Clinical Research delivery team, the “Clinical Research Network”.
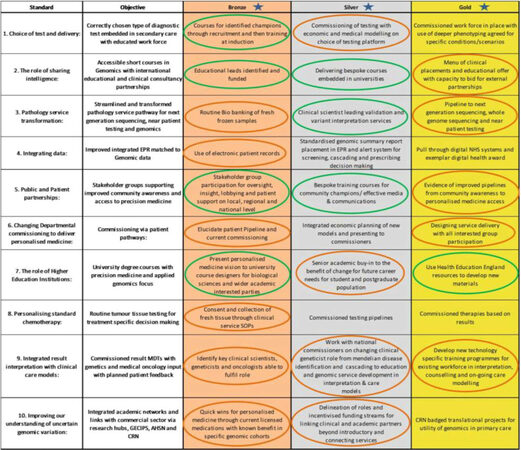
Patient involvement has been improved in four ways and (its) progress (is) assessed in Figure 2.
Figure 2. Assessment of genomic medicine progress (adapted from our select committee report submission[5])
Raising community awareness through classical and social media channels
Accounts of the “Angelina Jolie effect”[7] and the three sisters affected with breast cancer in less than a year and a half[8] were able to resonate with others to show the value of genetic testing and the impact it can have on health behaviour. With mainstreaming in mind, we have reached out to clinical colleagues in primary care and secondary care through talks, workshops, teaching sessions, webinars, triage aids, simple and streamlined referral systems, the media and the offer to embed staff in services. The pressures of acute hospital service delivery and the resulting financial and digital constraints, alongside delays in receiving results have at times made it difficult for colleagues to fully to embrace the potential benefits of genomic medicine.
Taking the message to the people through town centre mobile information centres and stalls at festivals
This has involved working with stakeholder charity groups to offer clinical advice and coordinate support for peer patient champions. With increasing availability of medical information online, our relationship with patients will evolve to include more sign-posting: providing opportunities for them to speak to other patients that have been through similar experiences. This is particularly relevant in Black and Minority Ethnic groups[9] where building trust through the use of community champions and the language used is important to ensure equity of access. In turn, this can aid the interpretation of rare variants and improve our understanding of the cause of diseases more prevalent in certain populations.
The development of academic partnerships
With the Clinical Research Network, the University of Leicester Precision Medicine Institute for personalised medicine, AstraZeneca and other pharmaceutical companies culminated in a conference held in March 2017[10]. This shaped a shared vision for genomic medicine and developed strategies for somatic testing of tumour tissue. We included patients in a regional stakeholder group to provide oversight to the clinical project and lobby commissioners and politicians (key decision makers). This approach helped link the National Institute of Health Research group, the Clinical Research Network and the 100,000 Genomes Project team locally. It informed policies on the involvement of seldom heard from groups. A similar national group including representatives from all of the regions has tackled issues such as readability of patient-facing literature, how patients wish to receive results and access to data.
Improving access to clinical services through simplified triage systems
The development of simple messages such as 321[11] in cancer genetics has facilitated communication and understanding of concepts in cancer triage (321: 3 affected relatives or more - across at least 2 generations and at least 1 diagnosed under the age of fifty). We developed visual aids for colleagues in mainstreamed specialities on entry criteria for the 100,000 Genomes Project. This was taken up nationally by Health Education England (HEE). In turn we have offered advice to the separate Czech, Norwegian and Finnish Genomic Strategy teams. The Finnish Genomic Medical board has encouraged us to continue to develop an improved assessment, triage, sign-post and communication system to involve the patient more seamlessly in genomic medicine. We aim to develop’ genomic responsive’ medical records to assist with prescribing when pharmacogenetics is incorporated into standard healthcare.
Education
HEE has a dedicated Genomics Education programme (GEP) which aims to help the delivery of the findings from the 100,000 Genomes Project into mainstream clinical practice by educating the NHS workforce[12,13]. In order to achieve this it has commissioned 7 UK universities to run a master’s level qualification in genomic medicine, with a syllabus designed for NHS professionals. This course can be undertaken as a full master’s degree (12 modules), a postgraduate diploma (8 modules), a postgraduate certificate (4 modules) or as individual continuing professional development modules in, for example, “pharmacogenetics and stratified medicine”. These courses have a set number of NHS commissioned places for eligible staff funded by GEP. It runs a module that directly uses data from the project and provides access to the GE research environment in a training-capacity only, but with the possibility of applying to the GeCIP to undergo research. HEE also funds fellowships in genomics within the NHS and genomics community, to try and stimulate active practitioners to use the 100,000 data for research that will inform their clinical practice. In addition, there is a free course on WGS and its implications, available online.
A roadmap designed in 2016 [Figure 2] to establish the standards of an effective genomic medicine service, including a bronze-silver-gold rating system of their implementation. Self-assessment can be made against these aims using a red-amber-green system for limited, significant and good progress. Our anticipated progress has been delayed somewhat by bioinformatic capacity challenges at the time of laboratory transformation and consolidation, delaying result giving and demonstrating patient benefit.
Consent
It is important that clinicians and patients have a frank and open discussion about the possibilities and limitations of genomic testing when consent is obtained. They must plan how the results are to be interpreted and communicated. There is understandably a desire to share positive news about families where mutations are identified that change management but this is still only the case for a minority of families at the present time. Patients need to be aware of the limitations of testing, particularly in chronic complex diseases. Individual patients need to have a careful discussion with their physician about how they wish to receive the results. Possible outcomes to be anticipated and considered included finding no clear pathogenic variant or conversely an unexpected incidental finding that may have future health implications.
The test directory
The NHS in England has established a directory of genetic tests available for patients with rare genetic disease or cancer. Tests will be offered in a standardised way across the nation, with clinical eligibility criteria stated in the directory. There will be a standardised consent form for genetic testing, with scope for patients to opt for involvement in future research projects and, when WGS is carried out, to receive additional incidental and potentially actionable findings. All of these developments build on lessons learnt in the 100,000 Genomes Project.
Tests in the directory may be ordered by a clinical geneticist or a specialist in secondary care. Although some of the processing of samples will need to be carried out locally in a timely fashion, the majority of the analysis will be carried out in one of a smaller number of large accredited laboratories that will be consolidated through the establishment of genomic laboratory hubs. Processing of blood, saliva or fresh tumour tissue to provide high quality samples for WGS is labour intensive and technically demanding, particularly when it is being established parallel to existing services with limited additional funding and current uncertainties about the future of local laboratories. This is compounded by a likely change to the role of pathology workforce to include more complex sample preparation to ensure the most informative parts of specimens are analysed and downstream bioinformatic interpretation. As is the case for many multi-step processes, any technical or motivational problem with patient identification, phenotyping, sample retrieval and processing, data analysis, clinical interpretation or communication to the patient could hinder the effective implementation of genomic medicine.
To aid interpretation of sequence variants in the future, we anticipate that the test results will be linked to a national database for phenotypes and genomic variants, but at the time of writing this is subject to confirmation. If busy secondary care clinicians are required to submit extensive phenotypic data when ordering a genetic test, we anticipate that they will need support to fit this into their existing patterns of work.
The test directory has been generated through combining the current UK Genetic Testing Network (UKGTN)[14] dossier of approved tests with PanelApp. Little detail on how this list was generated or evidence based with respect to variant call detection rate or economic modelling has been published, which makes critical analysis difficult.
Anecdotal patient stories from the 100,000 Genomes Project have been published about the impact of some test findings in individual cases. We look forward to seeing results collated from the different diseases and gene panels used to assess its health benefits in a more systematic way.
In lieu of arranging diagnostic testing in sporadic and non-syndromic disease and cancer, clinical geneticists will have vital additional roles in driving local innovation, education, supporting patient identification and variant interpretation. Patients with a complex family history, potentially syndromic disease, an identified variant, family dynamic/psychosocial challenges or a previous history of mental health problems will continue to need to be referred for genetic counselling. The focus will be to concentrate more on identifying clinically actionable variants in affected patients and then cascading results to at risk relatives. This contrasts with starting with the “worried well” approaching clinical genetics departments concerned about their family history, which is often currently the case.
Legacy
As the 100,000 Genomes Project finishes, it may be a challenge to maintain the wider engagement and dialogue around consent and data-sharing. The development of personalised medicine will target screening and specific treatments on those most likely to benefit rather than everyone in a population. Data storage and access are a controversial new currency in the digital age. Data about the health of an individual and his or her close relatives is highly sensitive information. There is a risk of increasing societal concerns about the direction that genomic medicine could take. We recommend that governments consider this an important aspect of transformational change otherwise technological advances may be unable to be successfully and appropriately implemented.
Challenges in clinical transformation
There is an often unspoken cynicism regarding whether new molecular diagnostics can improve healthcare in standard clinical settings when considering the benefits against other well-deserving healthcare priorities. This includes diagnostics for patients with acute medical problems or to guide treatment in patients with common chronic disease. This is partially due to two problems: firstly, a failure to recognize progress where present and secondly, a resistance to change in roles [Figure 3]. Failure to recognize the impact of molecular developments on patient outcomes when such innovations move from research into routine care is common. Education with evidence based economic modelling may be required here to demonstrate the value of such innovations. All diagnostics cost money while the benefits may come downstream in the patient pathway, and beyond. When economic modelling is used to plan healthcare the analysis must include the costs of companion diagnostics and any impact on any at risk relatives.
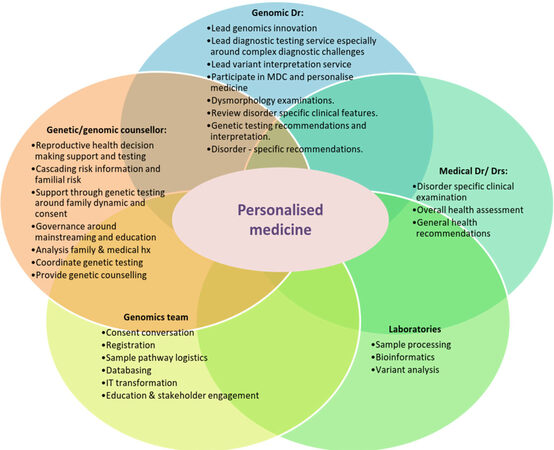
Figure 3. Roles of staff in genomic medicine. Proposed new staffing roles in the genomic medicine era
Secondly, change can be challenging and threatening to established clinicians, in spite of the simple fact that medicine changes as knowledge and technology grows. Change can be underpinned by evidence that the new intervention will solve an important problem and that it is worth changing priorities and effort alongside adapting or discarding tried and tested current practices.
Given competing pressures, implementation needs to be approached in three ways: (1) patient political drivers to demand change - these have been critical to a number of cancer therapies being introduced in the United Kingdom; (2) universal business templates that can be used at a local level to justify and explain how changes can be introduced beyond a national suggested guideline; and (3) crucially there must be clinical and managerial reasons to implement change beyond it being considered “the right thing to do”. This may include financial incentives or penalties. Perhaps greater focus would be achieved by linking the use of the diagnostic test with the ability to commission the relevant intervention. There is a case for restricting access to novel very expensive therapeutic agents to those most likely to benefit from such drugs. It seems likely that regulatory agencies, as well as funding bodies, may insist upon the use of companion diagnostic tests as stage in the patient pathway.
Ten recommendations
(1) Speak to stakeholders prior to starting a programme for operational design and buy-in; (2) build on existing expertise and strength using universal business templates and a combination of implementation incentives but also clear economic reasons for compliance with transformation - the “no diagnostic, no surgery” approach; (3) test and learn through rapid cycles based on simple criteria to inform implementation; (4) consider patient pathway economic modeling; (5) liaise with therapeutic companies around companion diagnostics; (6) consider impact on primary care of genomic results and how they are implemented; (7) be mindful of variable degrees of cynicism in primary and acute care around the importance of Mendelian disease in every day practice. Place an emphasis on approach to microbiology, cancer diagnostics and non-invasive disease; (8) ask patients about communication preferences around consent, re-contact and results; (9) don’t underestimate the power of patient stories but these anecdotal reports are not a substitute for academic rigour; and (10) be aware that genomic testing does not mean sequencing alone. It involves identification, registration and phenotyping, consent, sampling, processing, sequencing, interpretation, result giving, treatment and cascading.
Conclusion
Primary and secondary care physicians frequently claim that genetic testing can help a few patients with very rare disease but it fails to add diagnostic and prognostic power for common polygenic disorders. It is also asserted by many that the results rarely alter treatment or take too long to be available, so generic treatment has to be offered in the first instance. These concerns to date have been valid. Genomic testing has however proved to be of value in the diagnosis of rare inherited disease, particularly in presentations when one of a large number of candidate genes may underlie that individual patient’s phenotype. The 100,000 Genomes Project aims to change the way patients are diagnosed as well as how personalised treatments and cascading of risk information is carried out. For the lady mentioned at the beginning of the article, this means a fresh tumour sample being processed and analysed for somatic mutations that may be useful for designing treatment as well as identifying potentially significant variants that may be also present in the germline with wider implications for future cancer risk for the patient and/or their relatives. With routine tumour testing being instigated nationally and efficient family cascading, it is possible in time that the majority of families with mendelian susceptibility could be detected and thereby changing the role of the clinical geneticist and cementing links within medical oncology.
Declarations
AcknowledgmentsWe like to acknowledge the support of the East of England Stakeholder 100,000 Genomes Project partnership group.
Authors’ contributionsAll of the authors contributed intellectually and practically to this piece.
Availability of data and materialsNot applicable.
Financial support and sponsorshipNone.
Conflicts of interestThese are the individual views of the authors. Dr. Julian Barwell is also employed by the East Midlands Clinical Research Network and has received honoraria from AstraZeneca for running personalised medicine workshops.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2018.
REFERENCES
1. National Health Service. 100,000 Genomes Project Eligibility Wheels Library. Available from: https://www.genomicseducation.hee.nhs.uk/taught-courses/eligibility-wheels-library/. [Last accessed on 4 Sep 208].
2. SNOMED International. Leading healthcare terminology worldwide. Available from: https://www.snomed.org/participate. [Last accessed on 4 Sep 2018].
3. Genomics England PanelApp. A crowdsourcing tool to allow gene panels to be shared, downloaded, viewed and evaluated by the scientific community. Available from: https://panelapp.genomicsengland.co.uk/panels/. [Last accessed on 4 Sep 2018].
4. Genomics England. Working with participants, patient groups and the public. Available from: https://www.genomicsengland.co.uk/about-genomics-england/how-we-work/patient-and-public-involvement/. [Last accessed on Sep 2018].
5. UK Parliament. Genomics and genome-editing inquiry. Available from: https://www.parliament.uk/business/committees/committees-a-z/commons-select/science-and-technology-committee/inquiries/parliament-201/inquiry2/. [Last accessed on 4 Sep 2018].
6. Davies PDSC, Department of Health. Annual report of the chief medical officer 201: generation genome. Available from: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/31043/CMO_annual_report_generation_genome.pdf. [Last accessed on 4 Sep 2018].
7. Evans DG, Barwell J, Eccles DM, Collins A, Izatt L, Jacobs C, Donaldson A, Brady AF, Cuthbert A, Harrison R, Thomas S, Howell A, Miedzybrodzka Z, Murray A; FH02 Study Group. The Angelina Jolie effect: how high celebrity profile can have a major impact on provision of cancer related services. Breast Cancer Res 2014;16:442.
8. Sisters help launch cancer study aiming to end chemotherapy - BBC News. Available from: https://www.bbc.co.uk/news/health-35323066. [Last accessed on 4 Sep 2018].
9. Allford A, Qureshi N, Barwell J, Lewis C, Kai J. What hinders minority ethnic access to cancer genetics services and what may help? Eur J Hum Genet 2014;22:866-74.
10. UK Pharmacogenetics & Stratified Medicine Network. 4th annual open meeting. Available from: http://www.uk-pgx-stratmed.co.uk/images/stories/Publications/OnlineBrochure.pdf. [Last accessed on 4 Sep 2018].
11. National Cancer Research Insitute. Can the 321 triage system reduce the number of late referrals for familial cancer syndromes. Available from: http://abstracts.ncri.org.uk/abstract/can-the-321-triage-system-reduce-the-number-of-late-referrals-for-familial-cancer-syndromes/. [Last accessed on 4 Sep 2018].
12. Genomics Education Programme. Preparing for the future of healthcare. Available from: https://hee.nhs.uk/our-work/genomics-education. [Last accessed on 4 Sep 2018].
13. National Health Service. Master's in Genomic Medicine. Available from: https://www.genomicseducation.hee.nhs.uk/taught-courses/courses/masters-in-genomic-medicine/. [Last accessed on 4 Sep 2018].
14. National Health Service. UK Genetic Testing Network. Available from: https://ukgtn.nhs.uk/find-a-test/. [Last accessed on 4 Sep 2018].
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Barwell JG, O’Sullivan RBG, Mansbridge LK, Lowry JM, Dorkins HR. Challenges in implementing genomic medicine: the 100,000 Genomes Project. J Transl Genet Genom 2018;2:13. http://dx.doi.org/10.20517/jtgg.2018.17
AMA Style
Barwell JG, O’Sullivan RBG, Mansbridge LK, Lowry JM, Dorkins HR. Challenges in implementing genomic medicine: the 100,000 Genomes Project. Journal of Translational Genetics and Genomics. 2018; 2: 13. http://dx.doi.org/10.20517/jtgg.2018.17
Chicago/Turabian Style
Barwell, Julian G., Rory B.G. O’Sullivan, Laura K. Mansbridge, Joanna M. Lowry, Huw R. Dorkins. 2018. "Challenges in implementing genomic medicine: the 100,000 Genomes Project" Journal of Translational Genetics and Genomics. 2: 13. http://dx.doi.org/10.20517/jtgg.2018.17
ACS Style
Barwell, JG.; O’Sullivan RBG.; Mansbridge LK.; Lowry JM.; Dorkins HR. Challenges in implementing genomic medicine: the 100,000 Genomes Project. J. Transl. Genet. Genom. 2018, 2, 13. http://dx.doi.org/10.20517/jtgg.2018.17
About This Article
Special Issue
Copyright
Data & Comments
Data
 Cite This Article 88 clicks
Cite This Article 88 clicks













Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.